RxPlorer™ – Molecular Dynamics-Driven Metastate Discovery Screening: Eotaxin-1 as a Case Study
Introduction
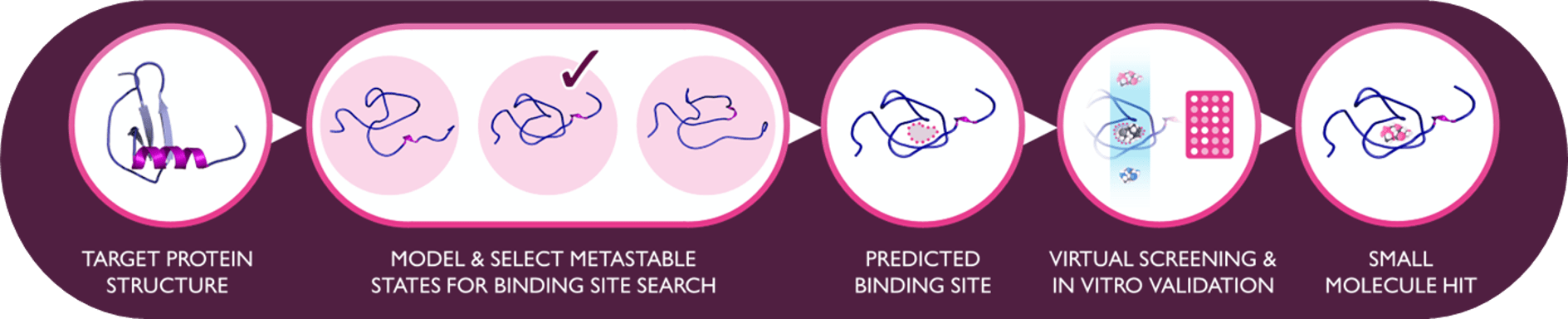
Proteins exist in dynamic conformational landscapes, but conventional structural biology techniques such as X-ray crystallography capture only a single low free energy conformation. As a result, many protein targets appear “undruggable” because whilst binding pockets are absent in such captured static structures, they can exist in low energy states despite being transient. Moreover, such transient low energy protein states can be structurally very distinct from wild type conformations, for example intermediates formed during folding pathways.
RxCelerate’s new approach, RxPlorer™, addresses this ability to identify distant-to-native but relevant protein binding sites by using a combination of accelerated long‑timescale molecular dynamics (MD) techniques to map a protein’s conformational space and identify metastable states. This novel method assigns likelihood to the existence of such states and relevance to conformational energy sampling. A hotspot‑mapping approach integrates this method to identify high‑confidence binding sites in such conformations, enabling rapid, structure‑based design and large‑scale virtual screening.
By revealing cryptic pockets and folding intermediates, RxPlorer™ facilitates rapid discovery campaigns where traditional methods fail – particularly for targets historically labelled “undruggable.” This method has been exemplified here on Eotaxin-1 (CCL11), a chemokine with no known small molecule binders. This case study identified metastable states and first-in-class small molecule modulators which may provide an alternative route to asthmas treatment instead of CCR3 (Eotaxin-1 receptor) antagonism, a long-standing target for the pharmaceutical industry.
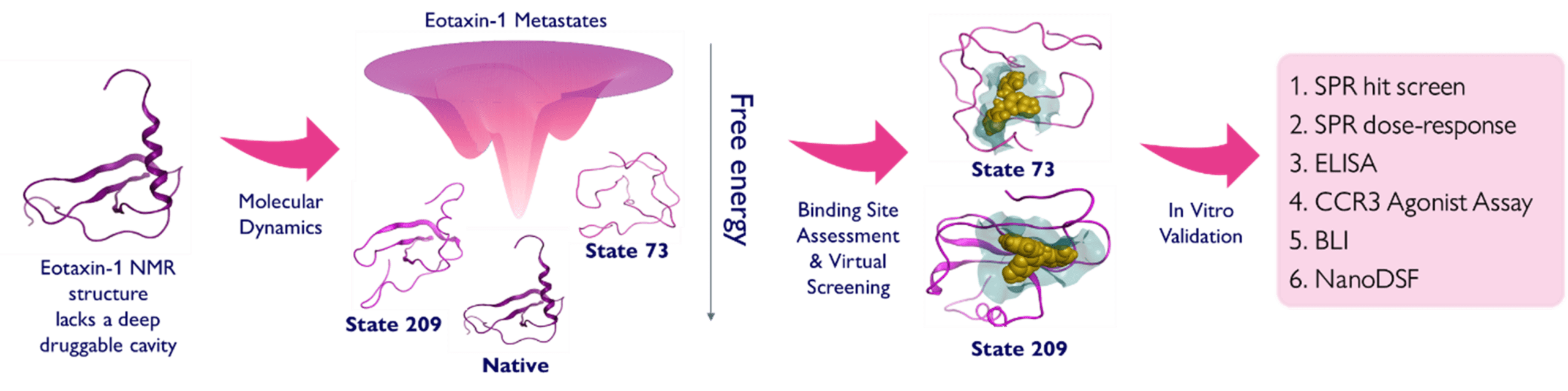
RxPlorer determines metastable states of Eotaxin-1 for structure based virtual screening
Long‑timescale accelerated MD simulations revealed two key metastable conformations of Eotaxin‑1: States 73 and 209, each exposing different cryptic, accessible pockets that are absent in the native structure. Both states are conformationally distant from the native form (backbone RMSD to native: State 73 = 0.78 nm; State 209 = 0.67 nm) yet are predicted to be transiently stable. These metastable conformations served as the structural basis for large‑scale virtual screening. As a comparator, virtual screening was also performed on the native state. 11 million compounds were screened separately against each state. 34 compounds in total were selected for in vitro assessment.

1. SPR Screening of 34 molecules identified hits for both metastates
Compounds were injected over captured Eotaxin‑1 for 60 s, generating molecular‑weight‑adjusted, reference‑subtracted responses for 34 molecules at 100 µM (SYN001–034). This initial screen identified binders across both metastable conformations predicted by RxPlorer™. Importantly, no hits were identified that were predicted to bind the native conformation of Eotaxin‑1.
A follow‑up dose‑response SPR assay confirmed target engagement for molecules that produced strong signals in the initial screen. Four compounds showing clear concentration‑dependent responses were selected for further characterisation.
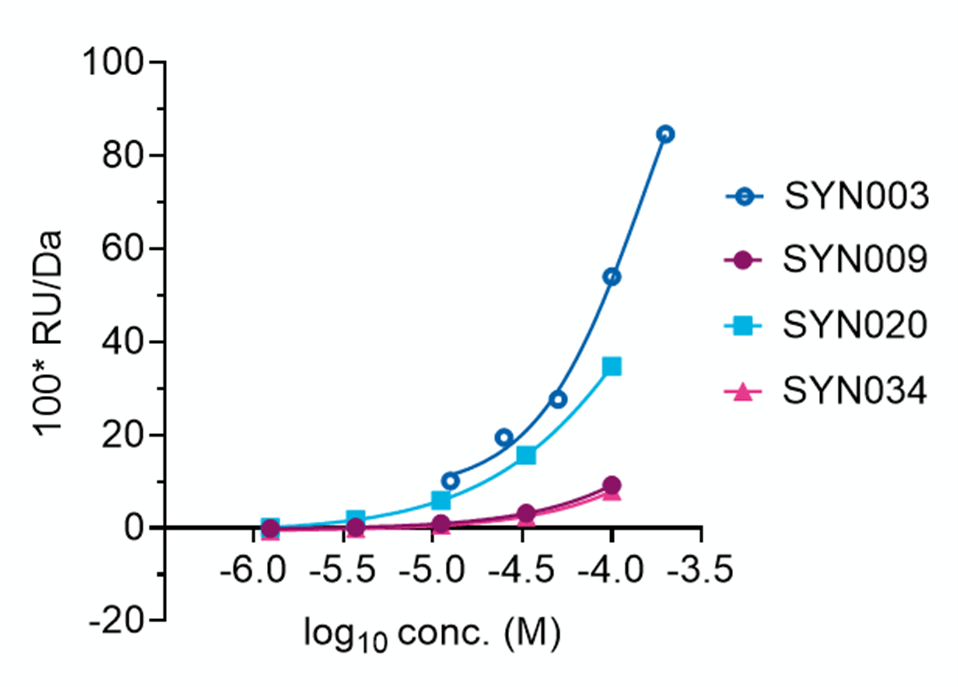
Molecular Weight Adjusted, Reference Subtracted binding (SPR)
2. ELISA data indicates molecules interfere with antibody epitope recognition on Eotaxin-1
An ELISA assay confirmed structural engagement: compounds SYN020 and SYN034 at 100 μM (SYN003 showed similar effects at 200 μM) inhibited antibody recognition of recombinant Eotaxin‑1. This indicates small molecule binding may induce conformational changes in Eotaxin-1, which occlude its antibody recognition epitope.
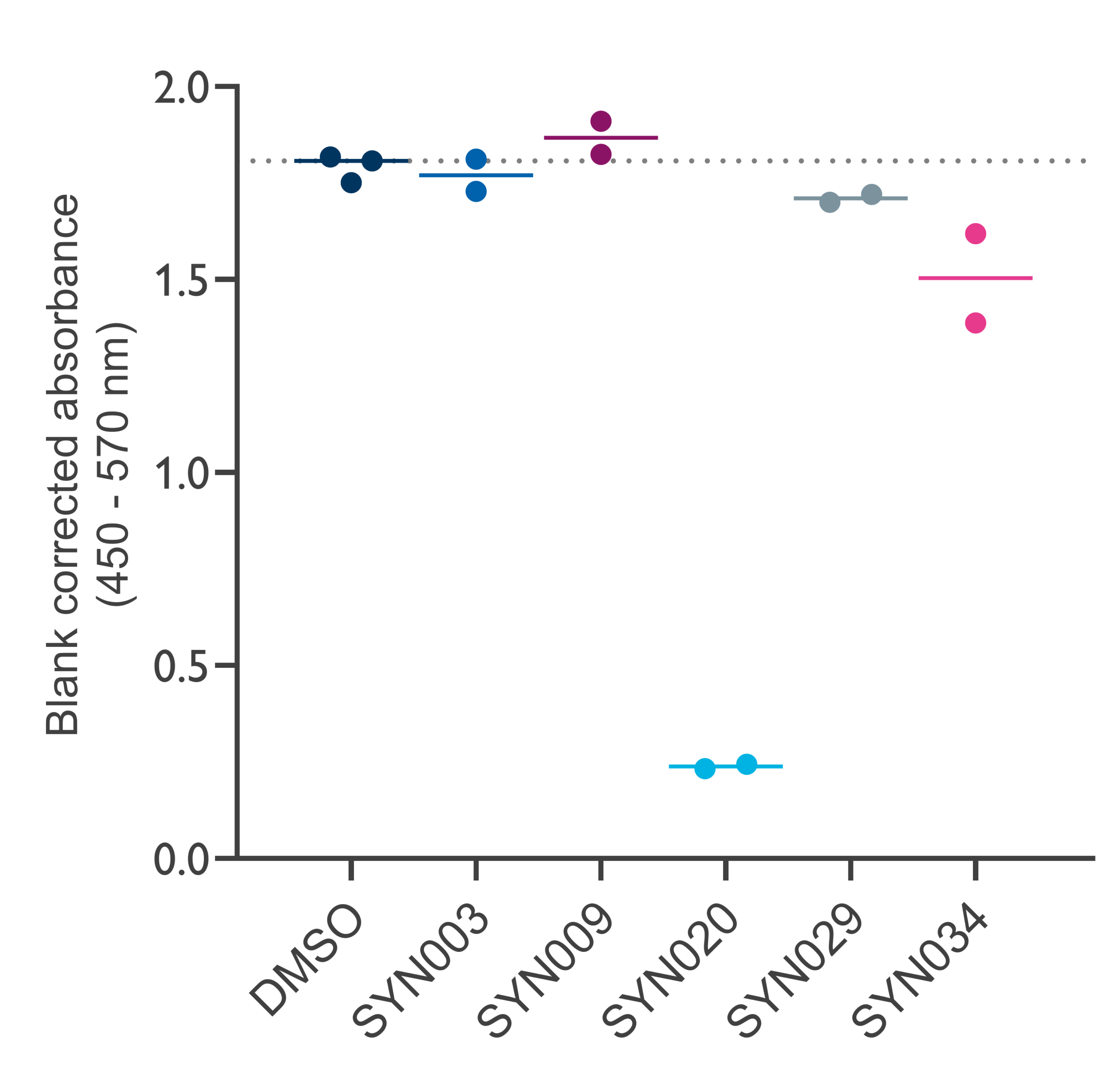
20 nM Eotaxin-1 binding to anti-Eotaxin-1 mAb on incubation with 100 μM test compound.
3. Inhibition of CCR3 signalling observed upon incubation of hit series with Eotaxin-1
A CCR3 Ca²⁺ agonist assay confirmed functional activity, with SYN003 showing the strongest reduction in signalling when incubated with Eotaxin‑1, demonstrating partial antagonism. Compounds in the absence of Eotaxin-1 had no effect on CCR3 signalling (data not shown).

Compounds with 10 nM Eotaxin-1 on CCR3 (agonist effect)
4. Extension of SAR with new molecules using BLI orthogonally indicates ligand-dependant conformational changes
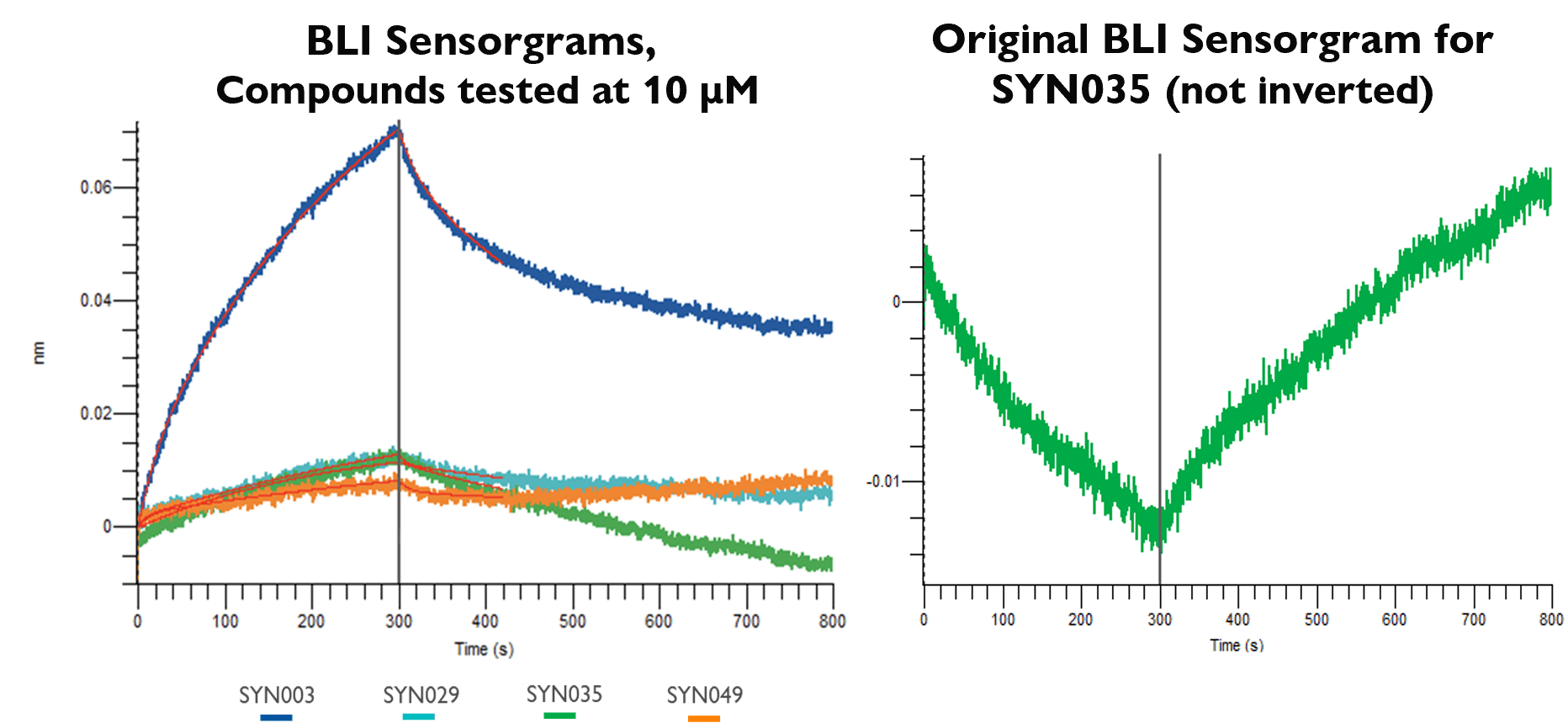
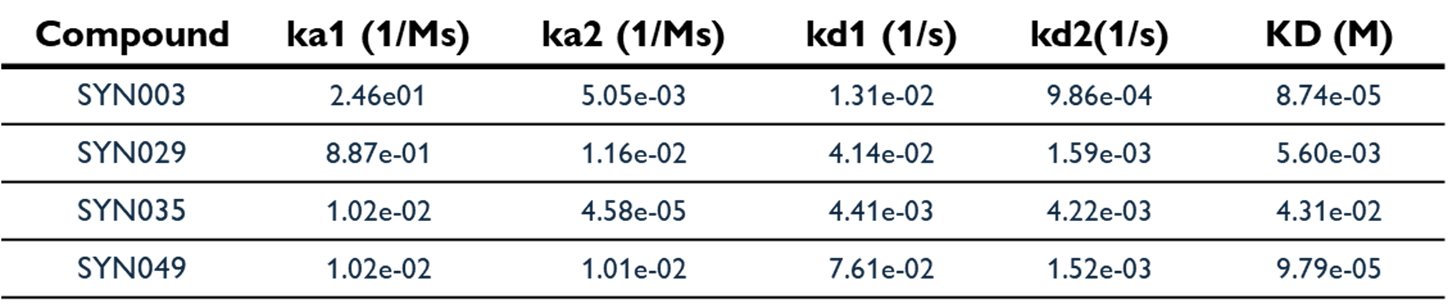
To explore the structure-activity relationship around the initial hits, SPR-screening of 25 additional analogues based on SYN003/009/020 was performed (data not shown). Further BLI assessment of lead compounds showed engagement, with clear two‑state kinetics consistent with a hypothesis of protein conformational change induced by ligand binding. SYN035 displayed complex kinetics with a negative response upon association that returned to baseline during dissociation. This is in line with the recent findings of FitzGerald et al. (2025) that suggest this profile is indicative of conformational change induced binding mechanisms. To report KD for SYN035, the sensorgram was inverted, which may have led to an apparent reduction in affinity*.
FitzGerald et al., RSC Chem. Biol., 2025,6, 1625-1639
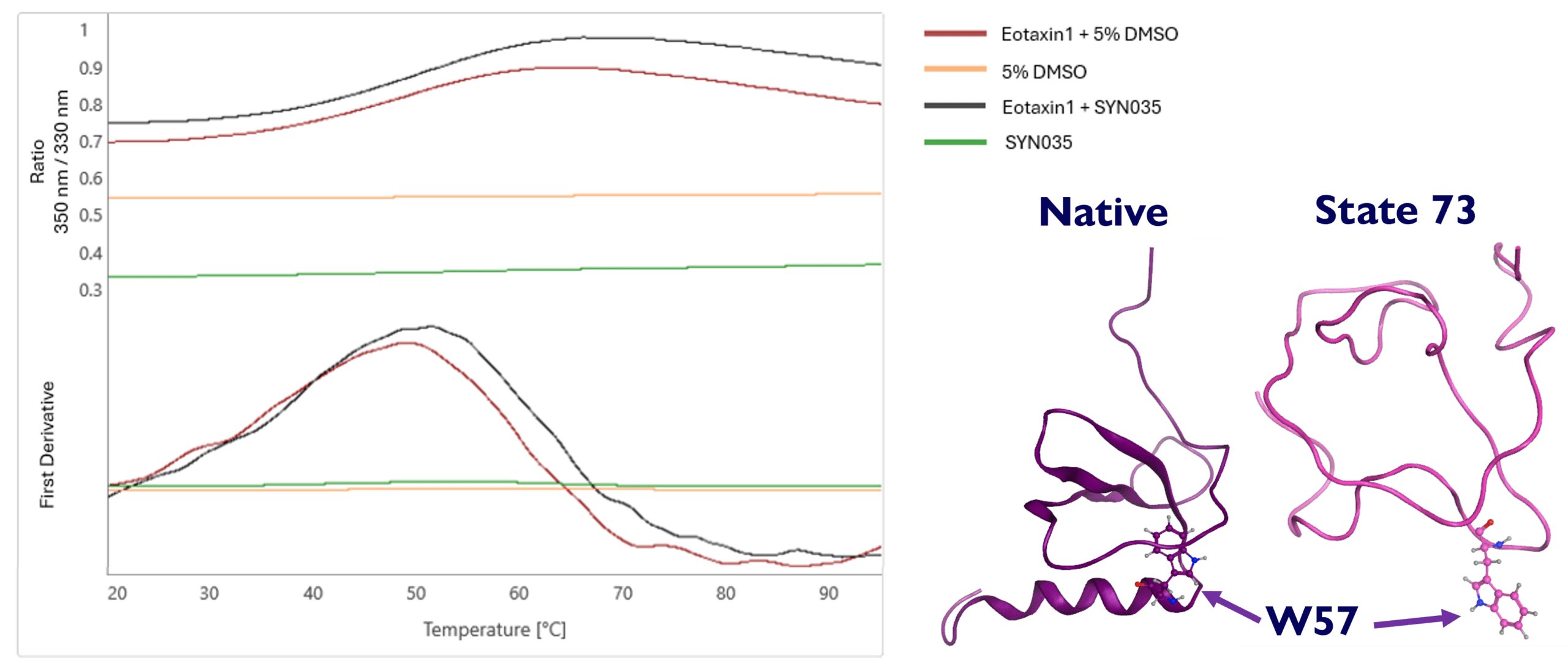
5. NanoDSF data indicates exposure of single Trp57 on Eotaxin-1
NanoDSF trace shows compound SYN035 increased thermal stability (+1.8 °C ΔTm to apo) and raised the 350/330 nm fluorescence ratio. This is predicted to be due to the change in solvent exposure of Eotaxin‑1’s single tryptophan (Trp57), which is buried in the native state. This tryptophan is exposed in the predicted structure of metastable state 73 when bound to SYN035. Hence, SYN035 binds to a conformation of Eotaxin-1 that is different to the native state; potentially a conformational state that is structurally close to state 73.
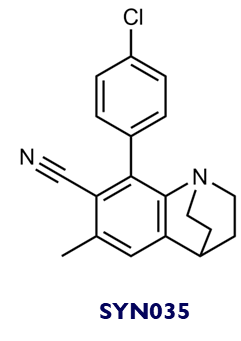
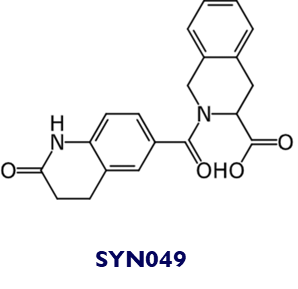
Chemical structure of SYN035 and SYN049: SYN035 and SYN049 (and their analogues) were identified as direct binders. Three different chemotype families were discovered in this campaign.
Summary
RxPlorer™ identified two druggable metastates of Eotaxin‑1, a previously unliganded chemokine, and enabled a focused virtual screen that yielded experimentally confirmed binders. SPR established direct target engagement, and subsequent assays demonstrated functional antagonism and conformational-dependent binding for particular compounds . These results show that metastate-guided screening reliably produces drug-like, functionally validated hits even for targets deemed “undruggable”. The RxPlorer™ platform enables the rapid discovery of novel hit series for challenging targets, expanding the chemical space available for drug design and accelerating drug discovery.
RxCelerate is the only organisation to offer metastate discovery as a dedicated service. Tell us about your challenging target and our scientists can begin work immediately, moving from modelling to in-house design, ordering and testing within two months.
RxPlorer™ launched in March 2026, and is currently available to clients through an early adopter programme. RxCelerate is inviting organisations seeking rapid discovery of novel hits for challenging targets to get in touch.
